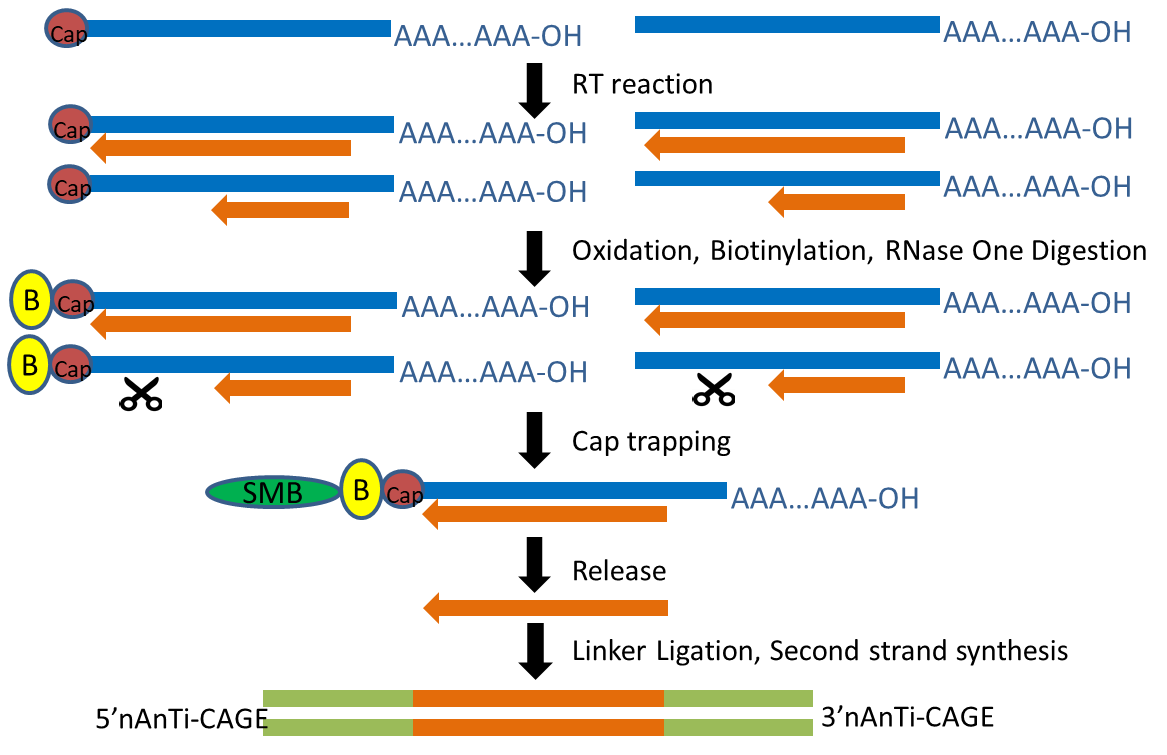
Comparison of CAGE and RNA-seq transcriptome profiling using clonally amplified and single-molecule next-generation sequencing | RNA-Seq Blog
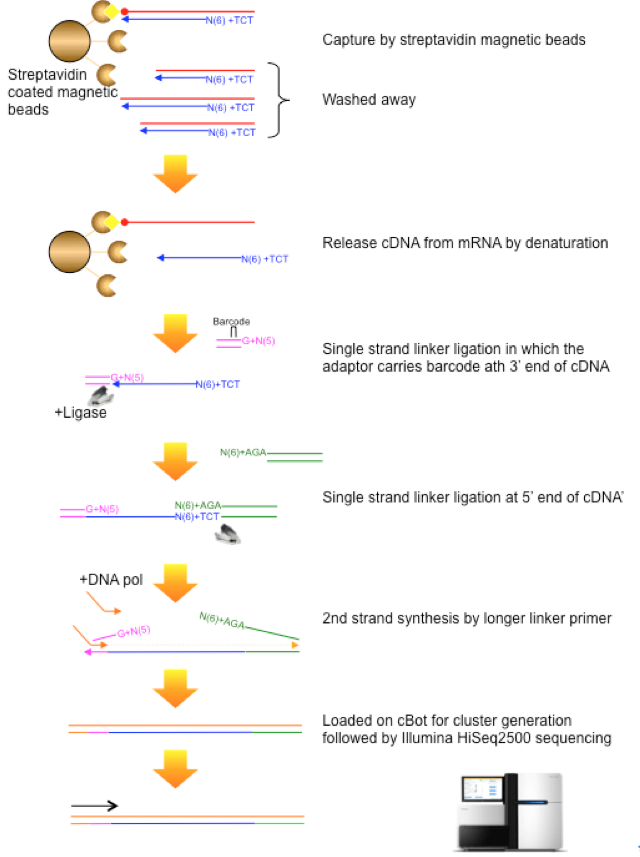
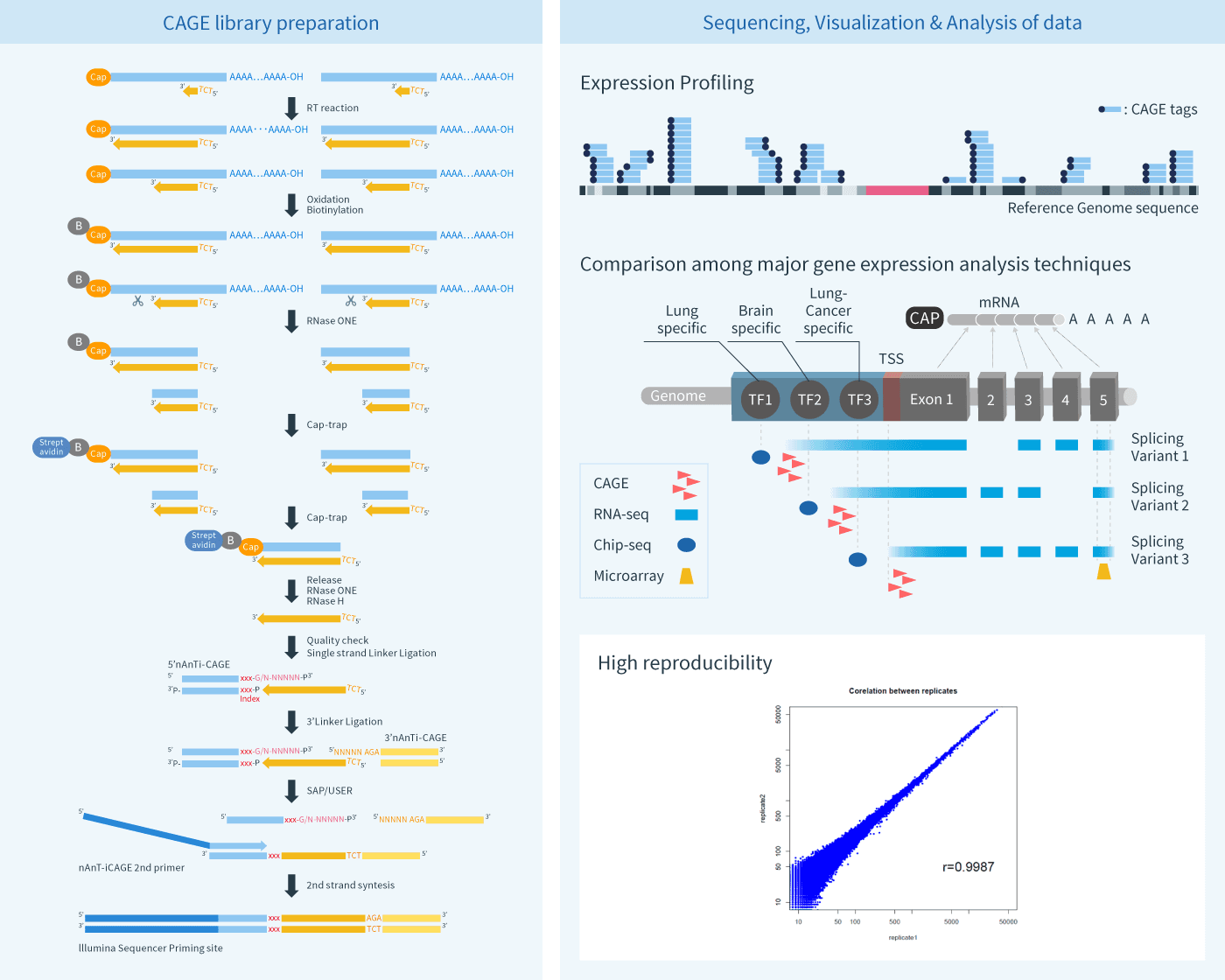
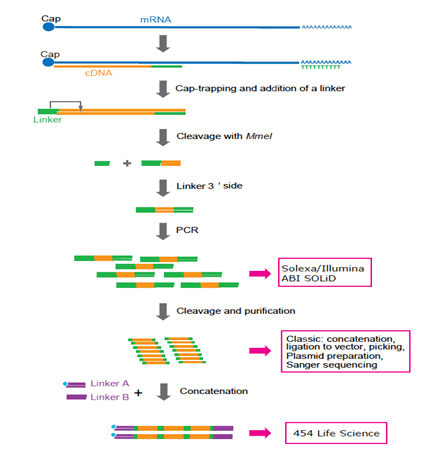
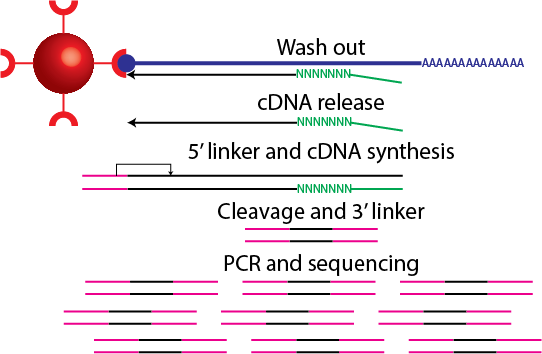
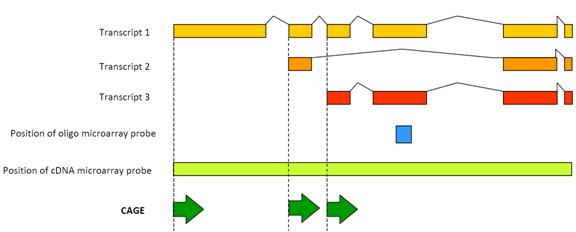
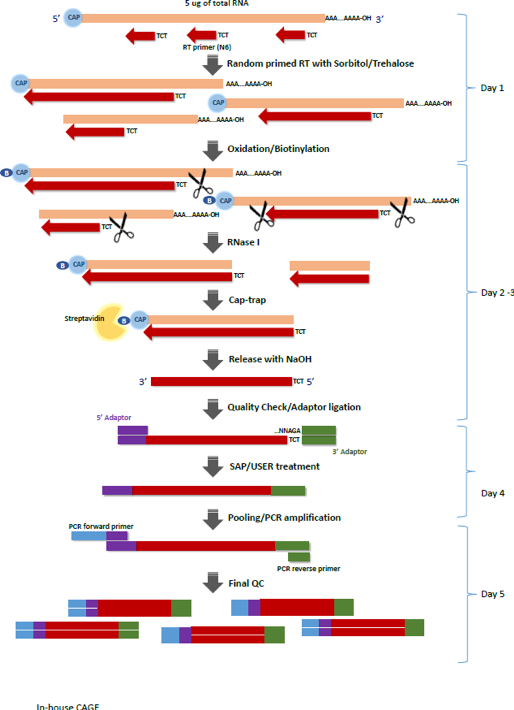
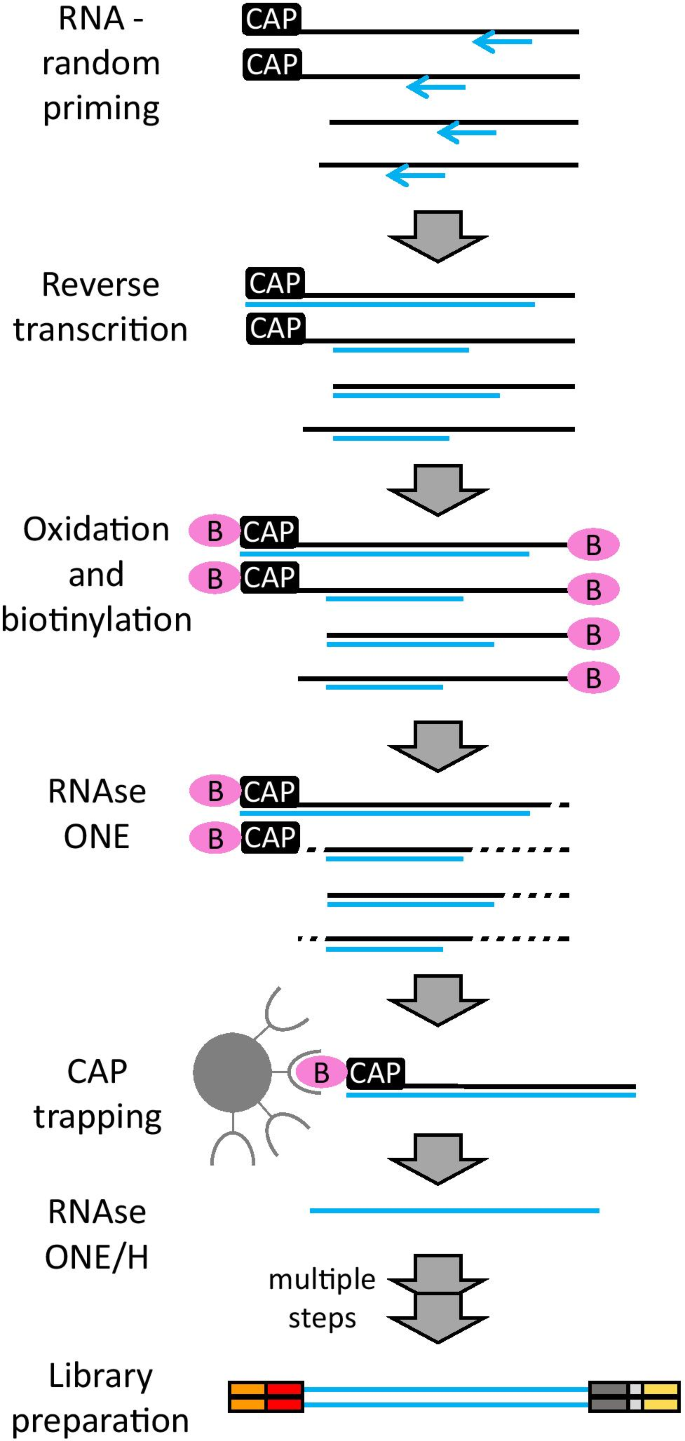
CAGE : A method for genome-wide identification of transcription start sites | 理化学研究所 ライフサイエンス技術基盤研究センター
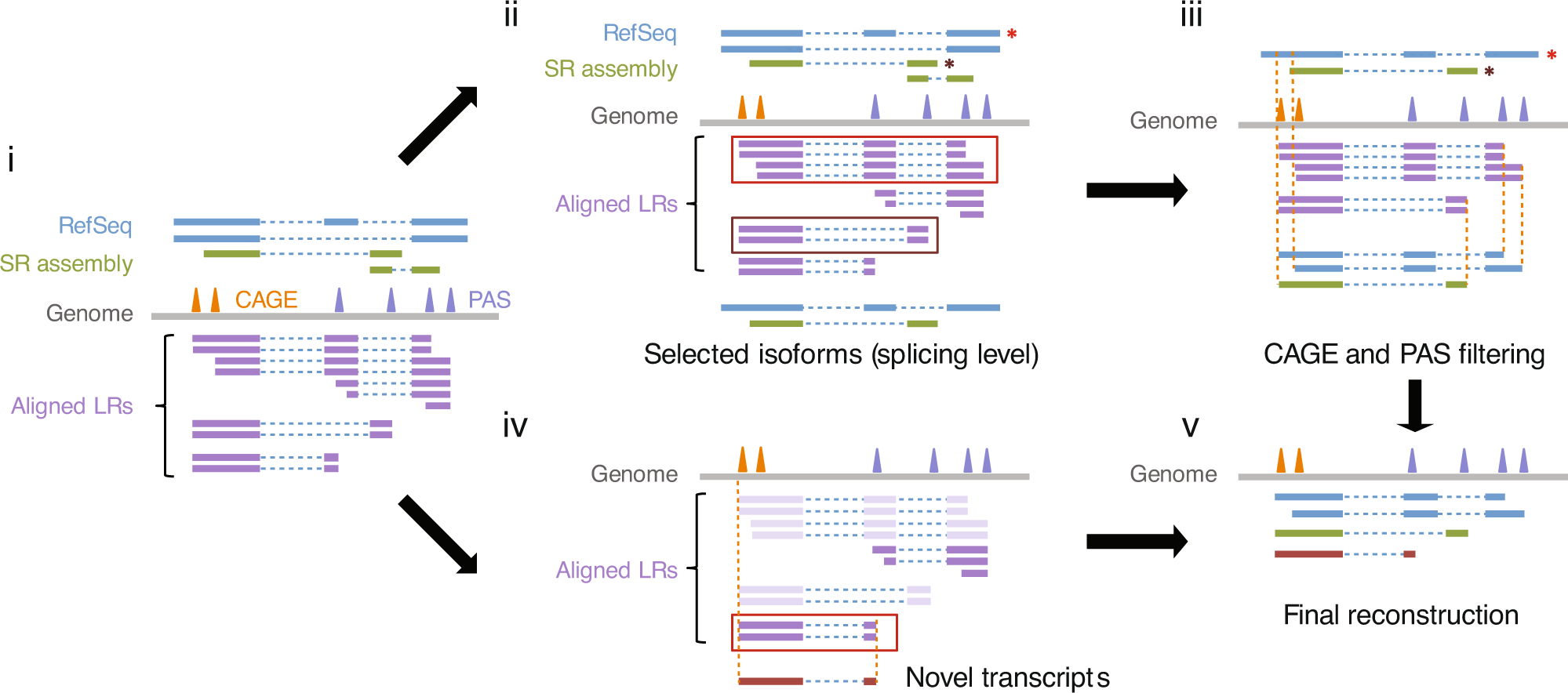
Single-molecule long-read sequencing reveals a conserved intact long RNA profile in sperm | Nature Communications

CAGE-sequencing identifies genome-wide activation of non-annotated TSSs... | Download Scientific Diagram
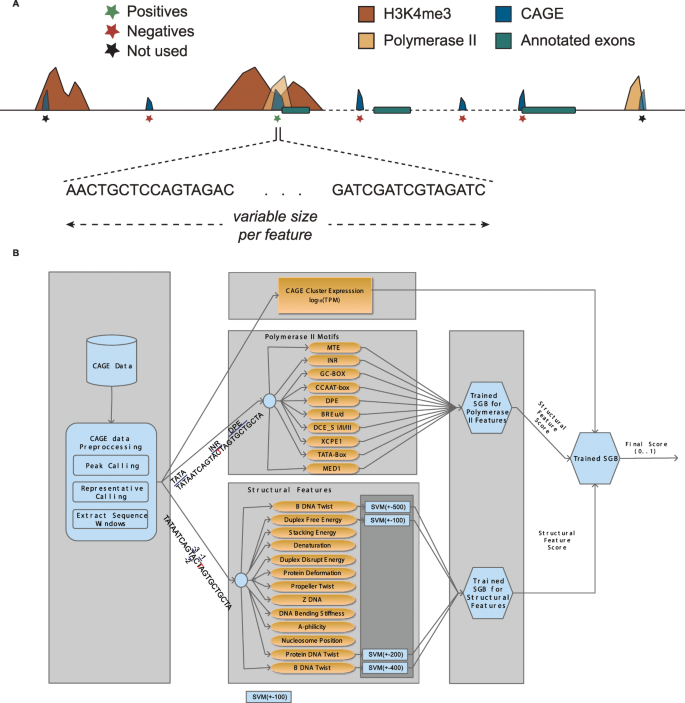
Solving the transcription start site identification problem with ADAPT-CAGE: a Machine Learning algorithm for the analysis of CAGE data | Scientific Reports

Figure 1 from CAGEr: an R package for CAGE (Cap Analysis of Gene Expression) data analysis and promoterome mining | Semantic Scholar
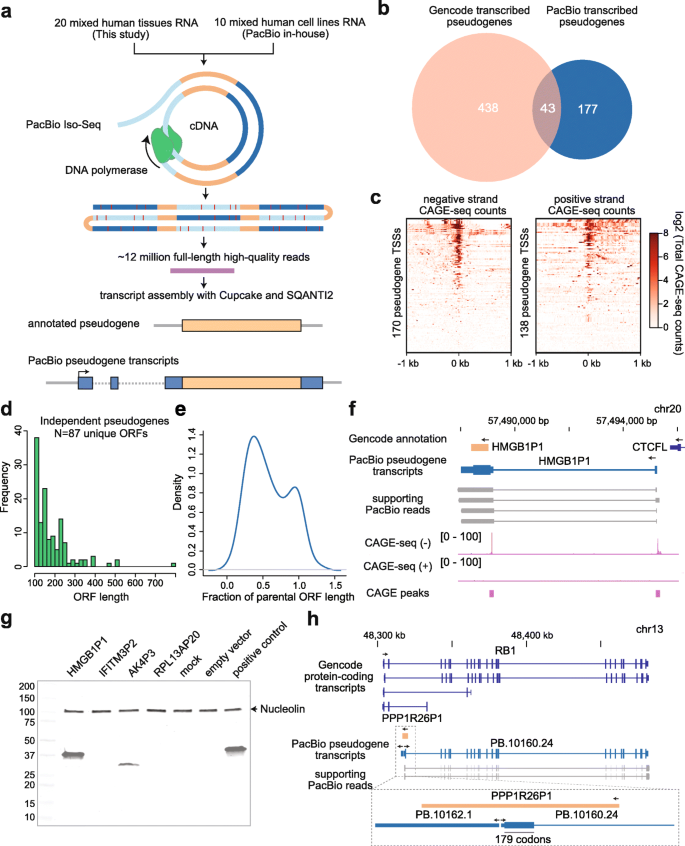
Long-read cDNA sequencing identifies functional pseudogenes in the human transcriptome | Genome Biology | Full Text

Cap analysis of gene expression (CAGE) sequencing reveals alternative promoter usage in complex disease | bioRxiv

piPipes – a set of pipelines for piRNA and transposon analysis via small RNA-seq, RNA-seq, degradome- and CAGE-seq, ChIP-seq and genomic DNA sequencing | RNA-Seq Blog

Key steps in the bioinformatic workflow for analysing CAGE sequencing... | Download Scientific Diagram